1.7.2.3. PitzDaily¶
1.7.2.3.1. Run Case¶
[5]:
import sys, time
import numpy as np
seed = 1435
np.random.seed(seed)
# import problem classes
import Exeter_CFD_Problems as TestProblems
verbose = False
sys.argv = sys.argv[:1]
print('Demonstration of the PitzDialy test problem.')
# set up directories.
settings = {
'source_case': 'Exeter_CFD_Problems/data/PitzDaily/case_fine/',
'case_path': 'Exeter_CFD_Problems/data/PitzDaily/case_single/',
'boundary_files': ['Exeter_CFD_Problems/data/PitzDaily/boundary.csv'],
'fixed_points_files': ['Exeter_CFD_Problems/data/PitzDaily/fixed.csv']
}
# instantiate the problem object
prob = TestProblems.PitzDaily(settings)
# get the lower and upper bounds
lb, ub = prob.get_decision_boundary()
# generate random solutions satisfying the lower and upper bounds.
x = np.array([ 1.54630729e-01, -4.99600344e-02, 1.38160660e-01, -4.99666603e-02,
2.28714652e-04, -3.24888531e-04, 2.74147357e-01, -1.88668377e-02,
6.62354130e-04, 1.86793810e-04])
#rand_x = []
#for i in range(x.shape[0]):
# if prob.constraint(x[i]): # check to see if the random solution is valid
# rand_x.append(x[i])
# evaluate for a solution
print('Number of control points: ', prob.n_control)
print('Decision vector: ', x)
print('Running simulation ...')
start = time.time()
res = prob.evaluate(x, verbose=verbose)
print('Objective function value:', res)
print('Time taken:', time.time()-start, ' seconds.')
Demonstration of the PitzDialy test problem.
Number of control points: [5]
Decision vector: [ 1.54630729e-01 -4.99600344e-02 1.38160660e-01 -4.99666603e-02
2.28714652e-04 -3.24888531e-04 2.74147357e-01 -1.88668377e-02
6.62354130e-04 1.86793810e-04]
Running simulation ...
Objective function value: 0.0799160834752
Time taken: 19.376554250717163 seconds.
[2]:
print(max(lb),min(lb))
-0.01 -0.05
[3]:
print(max(ub),min(ub))
0.287397 0.014
[4]:
x
[4]:
array([[ 5.78530692e-02, -3.31068377e-02, 5.50604920e-02, ...,
-4.71955674e-02, 1.88146599e-01, -1.32952546e-02],
[-5.67533117e-03, -7.48384848e-03, 1.38897278e-01, ...,
-3.25892854e-02, 2.22551915e-02, -7.50087530e-03],
[ 2.71721553e-02, -4.34923342e-02, 2.63459670e-01, ...,
-1.55789562e-02, 7.12139945e-02, -1.24998786e-02],
...,
[ 2.62482277e-01, 8.32653794e-03, 1.36335014e-01, ...,
-6.39175269e-03, 1.56653572e-01, -2.76899372e-02],
[ 2.32482592e-01, -2.66450403e-02, 1.01493483e-01, ...,
-2.29677976e-02, -8.32203134e-03, 1.86693078e-04],
[ 2.35917506e-02, -1.43564186e-02, -9.60306583e-03, ...,
-3.61561063e-02, 2.43942977e-01, -2.07163078e-03]])
1.7.2.3.2. Post Processing¶
[5]:
! ls
Exeter_CFD_Problems plotHeatExchanger.py
figures plotHeatExchangerTemperature.py
HeatExchanger.ipynb plotKaplanDuct.py
KaplanDuct.ipynb plotPitzDailyGrid.py
LICENSE plotPitzDailyStreamlines.py
log plotPitzDailyVelocity.py
log2 PlyParser_FoamFileParser_parsetab.py
log3 __pycache__
log4 README.md
PitzDaily.ipynb sample_script.py
PitzDailyPoints.ipynb U_centerplane.vtk
[6]:
! postProcess -case Exeter_CFD_Problems/data/PitzDaily/case_single -func sample > log 2>&1
[7]:
# I have run this case from cases_fine to see what the results are for non_optimised case
! postProcess -case Exeter_CFD_Problems/data/PitzDaily/case_non_optimised -func sample > log2 2>&1
[6]:
import warnings; warnings.simplefilter('ignore')
[15]:
%matplotlib inline
[16]:
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from turbulucid import *
matplotlib.rcParams['figure.figsize'] = (15, 10)
[17]:
matplotlib.rcParams['axes.labelsize'] = 15
matplotlib.rcParams['xtick.labelsize'] = 15
matplotlib.rcParams['ytick.labelsize'] = 15
matplotlib.rcParams['text.usetex'] = True
[8]:
from os.path import join
import turbulucid
import importlib
importlib.reload(turbulucid) # have to reload module if changes are made
case = Case(join("./Exeter_CFD_Problems/data/PitzDaily/", \
"case_single","postProcessing","sample", "0", "U_nearWall.vtk"),pointData=True)
[13]:
print(case.fields)
['U']
[14]:
case.xlim[0]
[14]:
-0.025776666930589998
[15]:
case.ylim[0]
[15]:
-0.026246666349433335
[16]:
case.xlim[1]
[16]:
0.29517665818919003
[17]:
max(case.cellCentres[0])
[17]:
0.00020892665
[18]:
case.cellCentres
[18]:
array([[-1.9027714e-02, 2.0892665e-04],
[-1.9027714e-02, 1.0446333e-04],
[-2.0072391e-02, 3.1876573e-04],
...,
[ 2.6091382e-01, -1.9279622e-02],
[ 2.5608581e-01, -1.9622434e-02],
[ 2.5503725e-01, -1.9715633e-02]], dtype=float32)
[19]:
case['U']
[19]:
array([[ 9.8294477e+00, -8.0274665e-01, 5.5096077e-25],
[ 6.5214849e+00, -7.0288688e-01, 4.2253022e-25],
[ 9.9577675e+00, -2.6880601e-01, -3.0189468e-24],
...,
[ 1.3043121e+00, 1.0591840e-01, 0.0000000e+00],
[ 1.2289770e+00, 7.9705156e-02, -2.8791849e-19],
[ 1.2289770e+00, 7.9705156e-02, -2.8791849e-19]], dtype=float32)
[20]:
case["magU"] = np.linalg.norm(case["U"], axis=1)
[21]:
case["magU"]
[21]:
array([9.86217213, 6.55925417, 9.96139526, ..., 1.30860567, 1.2315588 ,
1.2315588 ])
[22]:
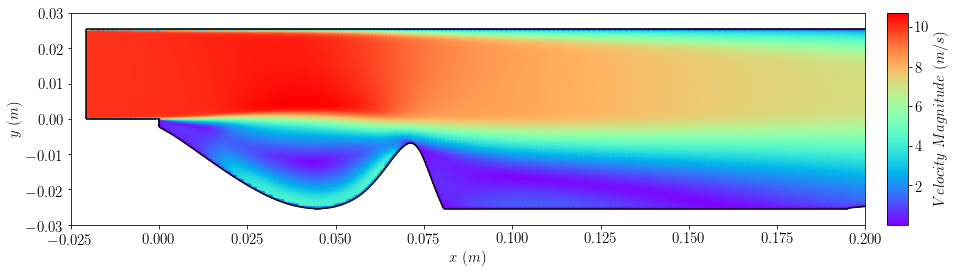
plot=plot_field(case, "magU", cmap='rainbow',colorbar=False)
plt.xlim([-0.025,0.2])
plt.ylim([-0.03,0.03])
plt.xlabel(r'$x \ (m)$')
plt.ylabel(r'$y \ (m)$')
bar=add_colorbar(plot, aspect=10, padFraction=1)
bar.set_label(r'$Velocity \ Magnitude \ (m/s)$')
[23]:
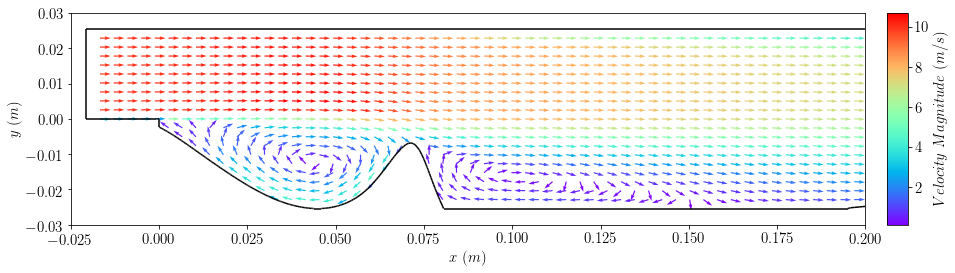
plot2=plot_vectors(case,'U', colorField='magU',cmap='rainbow', normalize=True, \
sampleByPlane=True,planeResolution=[20, 80], width=0.0015)
plt.xlim([-0.025,0.2])
plt.ylim([-0.03,0.03])
plt.xlabel(r'$x \ (m)$')
plt.ylabel(r'$y \ (m)$')
bar2=add_colorbar(plot2, aspect=10, padFraction=1)
bar2.set_label(r'$Velocity \ Magnitude \ (m/s)$')
[17]:
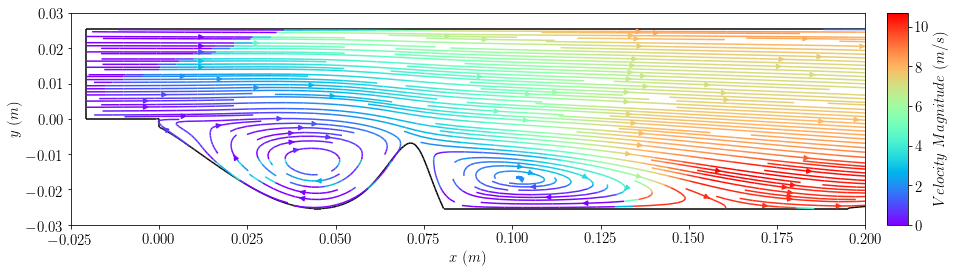
plot3=plot_streamlines(case, 'U',colorField='magU',cmap='rainbow',planeResolution=[100,100],density=2)
plt.xlim([-0.025,0.2])
plt.ylim([-0.03,0.03])
plt.xlabel(r'$x \ (m)$')
plt.ylabel(r'$y \ (m)$')
bar3=add_colorbar(plot3.lines, aspect=10, padFraction=1)
bar3.set_label(r'$Velocity \ Magnitude \ (m/s)$')
[9]:
case2 = Case(join("./Exeter_CFD_Problems/data/PitzDaily/", \
"case_non_optimised","postProcessing","sample", "1634", "U_nearWall.vtk"),pointData=True)
[37]:
case2["magU"] = np.linalg.norm(case2["U"], axis=1)
[22]:
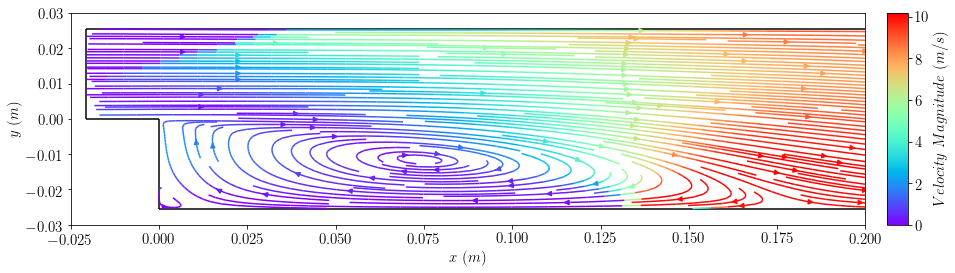
plot4=plot_streamlines(case2, 'U',colorField='magU',cmap='rainbow',planeResolution=[90,90],density=2)
plt.xlim([-0.025,0.2])
plt.ylim([-0.03,0.03])
plt.xlabel(r'$x \ (m)$')
plt.ylabel(r'$y \ (m)$')
bar4=add_colorbar(plot4.lines, aspect=10, padFraction=1)
bar4.set_label(r'$Velocity \ Magnitude \ (m/s)$')
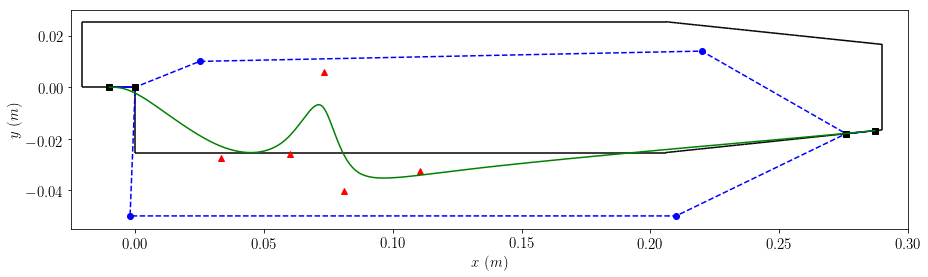
1.7.2.3.3. Plot Catmull-Clark Curve with Decision Vector, Boundary Points and Fixed Points¶
My makeshift way of extracting the decision vector, boundary points and fixed points: * for the decision vector I have extracted the values from rand_x * for the boundary points I have read in the file prob.domain_files * for the fixed points I have read in the file prob.fixed_points_files
Ideally I should have created functions to extract and order and plot these values instead of this, however it does seem to be correct
[11]:
# Partial re-run of the case to access values for plotting
import sys, time
import numpy as np
seed = 1435
np.random.seed(seed)
# import problem classes
import Exeter_CFD_Problems as TestProblems
verbose = False
sys.argv = sys.argv[:1]
print('Demonstration of the PitzDialy test problem.')
# set up directories.
settings = {
'source_case': 'Exeter_CFD_Problems/data/PitzDaily/case_fine/',
'case_path': 'Exeter_CFD_Problems/data/PitzDaily/case_single/',
'boundary_files': ['Exeter_CFD_Problems/data/PitzDaily/boundary.csv'],
'fixed_points_files': ['Exeter_CFD_Problems/data/PitzDaily/fixed.csv']
}
# instantiate the problem object
prob = TestProblems.PitzDaily(settings)
# get the lower and upper bounds
lb, ub = prob.get_decision_boundary()
# generate random solutions satisfying the lower and upper bounds.
x = np.random.random((1000, lb.shape[0])) * (ub - lb) + lb
rand_x = []
for i in range(x.shape[0]):
if prob.constraint(x[i]): # check to see if the random solution is valid
rand_x.append(x[i])
Demonstration of the PitzDialy test problem.
[29]:
boundaries=plot_boundaries(case2)
plt.xlim([-0.025,0.3])
plt.ylim([-0.055,0.03])
plt.xlabel(r'$x \ (m)$')
plt.ylabel(r'$y \ (m)$')
# decision vector
for i in range(0,10,2):
plt.plot(rand_x[0][i],rand_x[0][i+1], 'r^')
# boundary points
pts = [np.loadtxt(filename, delimiter=',') for filename in prob.domain_files]
x_boundary = []
y_boundary = []
for i in range(0,11,1):
x_boundary.append(pts[0][i][0])
y_boundary.append(pts[0][i][1])
plt.plot(x_boundary,y_boundary,'bo--')
#fixed_points
fixed_points = [list(np.loadtxt(filename, delimiter=',').astype(int))\
for filename in prob.fixed_points_files]
fixed=[fixed_points[0][0][0],fixed_points[0][0][1],fixed_points[0][1][0],fixed_points[0][1][1]]
x_fixed = []
y_fixed = []
for j in fixed:
x_fixed.append(pts[0][j][0])
y_fixed.append(pts[0][j][1])
plt.plot(x_fixed,y_fixed,'ks')
x_curve = []
y_curve = []
#catmull_clark curve
shape=prob.convert_decision_to_shape(rand_x[0]) # convert random array to ordered list
for control_points in shape:
data_points = TestProblems.data.support.catmull(control_points)
for i in range(prob.niter):
data_points = TestProblems.data.support.catmull(data_points)
for k in range(0,512):
x_curve.append(data_points[k][0])
y_curve.append(data_points[k][1])
plt.plot(x_curve,y_curve,'g-')
[29]:
[<matplotlib.lines.Line2D at 0x7fe2bc679be0>]
[27]:
len(pts[0])
[27]:
11
[22]:
pts #these are the boundary points [x, y] there are 11 of them
[22]:
[array([[-0.01 , 0. ],
[ 0. , 0. ],
[ 0.025 , 0.01 ],
[ 0.22 , 0.014 ],
[ 0.276 , -0.0180675],
[ 0.287397 , -0.0168678],
[ 0.276 , -0.0180666],
[ 0.21 , -0.05 ],
[-0.002 , -0.05 ],
[ 0. , 0. ],
[-0.01 , 0. ]])]
[23]:
fixed_points # these are the fixed points, the first two points, and points 4 and 5. They are grouped like
# this to control the gradient at these points
[23]:
[[array([0, 1]), array([4, 5])]]
Question: why are there only four fixed points, when there should be seven, because points 6, 9 and 10 are also fixed? Is this something to do with the gradient being controlled only by [0,1] and [4,5]? Or are they arbitarily chosen to be the ends of the polynomial only? i.e. I could have fixed [5,6] and [9,10] instead of [0,1] and [4,5]
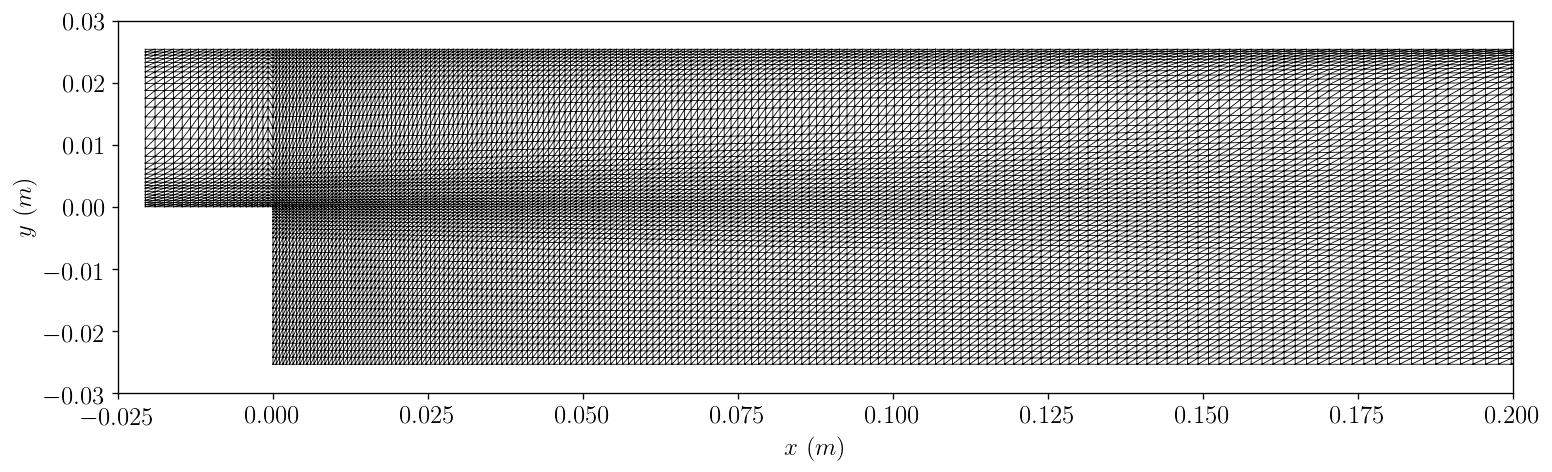
1.7.2.3.4. Triangulated Grids¶
[52]:
import plotPitzDailyGrid
import importlib
importlib.reload(plotPitzDailyGrid) # have to reload module if changes are made
%matplotlib inline
[53]:
! foamToVTK -case Exeter_CFD_Problems/data/PitzDaily/case_non_optimised > log3 2>&1
grid = \
'Exeter_CFD_Problems/data/PitzDaily/case_non_optimised/VTK/case_non_optimised_1634.vtk'
plotPitzDailyGrid.plot(grid)
[54]:
! foamToVTK -case Exeter_CFD_Problems/data/PitzDaily/case_single > log4 2>&1
grid_2 = \
'Exeter_CFD_Problems/data/PitzDaily/case_single/VTK/case_single_0.vtk'
plotPitzDailyGrid.plot(grid_2)
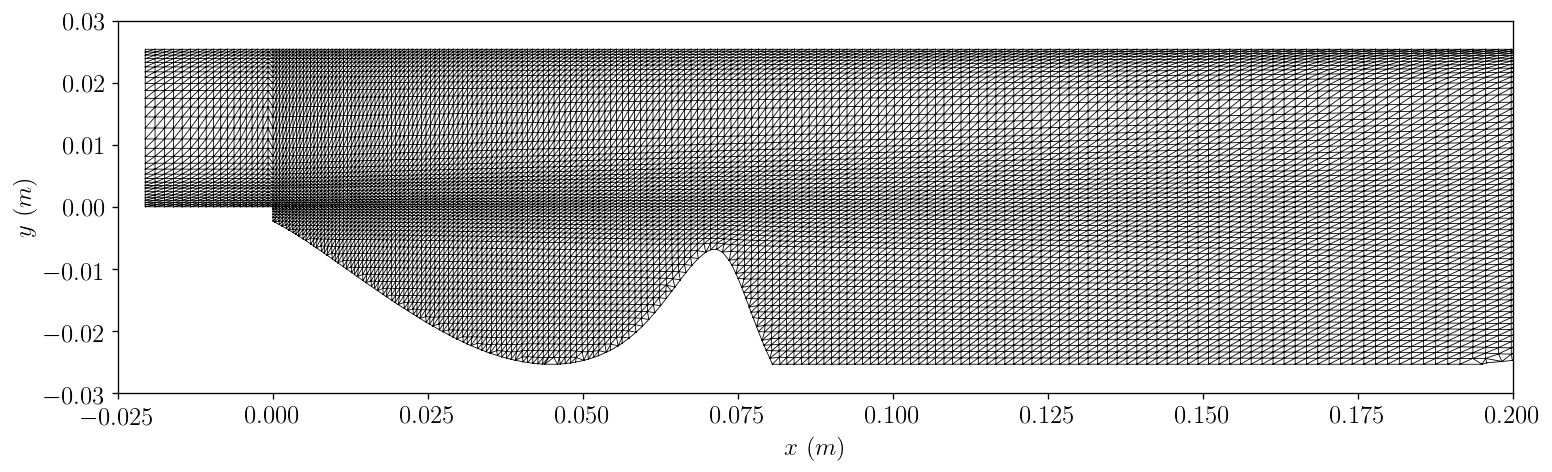
Question: Why is some of the Catmull-Clark polynomial ignored if y < -0.025m, even though it is within the permitted boundary?
[ ]: